MSA Advanced Scoring
Instructions
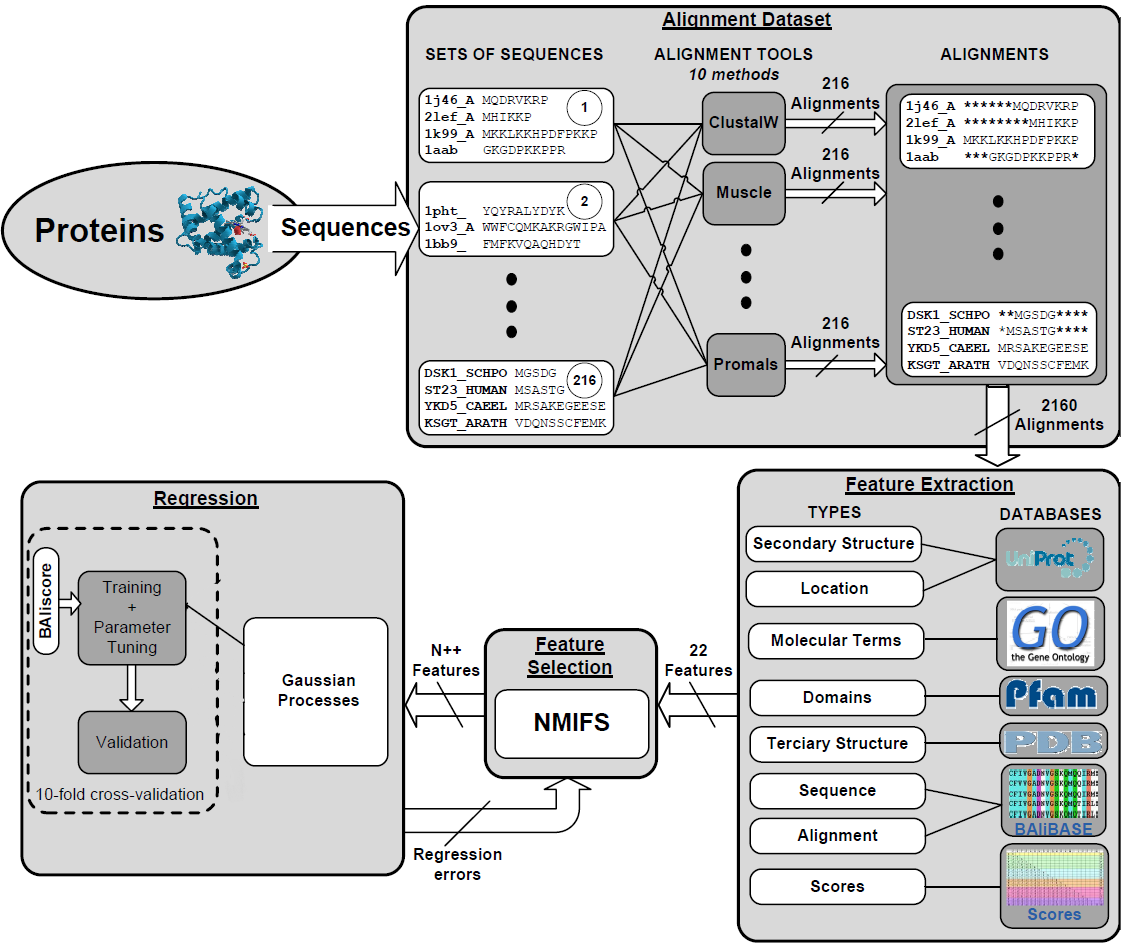
1. Intelligent score scheme to calculate the quality of multiple sequence alignments.
2. This score scheme is based on a Gaussian Process model trained by 15 biological features.
3. Select your alignment file (FASTA or MSF format) and click submit to obtain its score.
4. Sequence headers in this file should be valid PDB or Uniprot accessions.
5. Please, click "Run Example" to run the score scheme for this alignment file.